This notebook demonstrates blackjax.laplace_hmc, a sampler that integrates out
latent Gaussian variables via the Laplace approximation and runs HMC over the
marginal hyperparameter posterior.
Core idea. In a hierarchical model
phi ~ p(phi) # hyperparameters (low-dimensional)
theta ~ N(0, K(phi)) # latent variables (high-dimensional)
y ~ p(y | theta, phi) # likelihoodsampling the joint posterior is hard because the geometry changes dramatically with (“funnel” geometry). Laplace-HMC instead runs HMC over alone, using the Laplace approximation to analytically handle . The mode is found via L-BFGS, and the log-marginal gradient uses the implicit function theorem — no gradient unrolling through the optimizer.
This notebook demonstrates:
Efficiency on Neal’s Funnel — where Laplace HMC excels by marginalizing away the difficult geometry.
GP Regression and Accuracy — where the Laplace approximation is exact.
Comparison of Sampler Variants — benchmarking fixed/dynamic and standard/multinomial variants.
import time
import jax
import jax.numpy as jnp
import matplotlib.pyplot as plt
import numpy as np
import blackjax
import numpyro
import numpyro.distributions as dist
from numpyro.infer.util import initialize_model
# Reproducibility
from datetime import date
key = jax.random.key(int(date.today().strftime("%Y%m%d")))
plt.rcParams.update({"figure.dpi": 120, "font.size": 11})
def inference_loop(rng_key, kernel, initial_state, num_samples):
@jax.jit
def one_step(state, rng_key):
state, info = kernel(rng_key, state)
return state, (state, info)
keys = jax.random.split(rng_key, num_samples)
return jax.lax.scan(one_step, initial_state, keys)
n_chains_default = 8
def run_chains(rng_key, kernel, initial_state, num_samples, n_chains=n_chains_default):
def one_chain(key, state):
_, (states, info) = inference_loop(key, kernel, state, num_samples)
return states, info
keys = jax.random.split(rng_key, n_chains)
# Replicate initial_state if not already batched
leaf = jax.tree.leaves(initial_state)[0]
if leaf.ndim == 0 or leaf.shape[0] != n_chains:
initial_state = jax.tree.map(
lambda x: jnp.repeat(x[None, ...], n_chains, axis=0), initial_state
)
return jax.vmap(one_chain)(keys, initial_state)/opt/hostedtoolcache/Python/3.13.12/x64/lib/python3.13/site-packages/tqdm/auto.py:21: TqdmWarning: IProgress not found. Please update jupyter and ipywidgets. See https://ipywidgets.readthedocs.io/en/stable/user_install.html
from .autonotebook import tqdm as notebook_tqdm
Section 1: Neal’s Funnel (Speed and Efficiency)¶
Neal’s Funnel is a classic example of a “difficult” geometry for MCMC. The joint distribution is:
In the joint space, as becomes small, the conditional distribution of becomes very narrow, forcing HMC to take tiny steps. Laplace HMC marginalizes out analytically (the approximation is exact here!), leaving a simple Gaussian for .
n_theta = 100
def funnel_model(n_obs=100):
v = numpyro.sample("v", dist.Normal(0.0, 3.0))
with numpyro.plate("data", n_obs):
numpyro.sample("theta", dist.Normal(0.0, jnp.exp(v)))
init_key, key = jax.random.split(key)
_, potential_fn, _, _ = initialize_model(init_key, funnel_model, model_args=(n_theta,))
def laplace_log_joint(theta, v):
return -potential_fn({'theta': theta, 'v': v})
def nuts_log_joint(params):
return -potential_fn(params)Laplace HMC vs NUTS¶
# 1. Setup Laplace Marginal
laplace_funnel = blackjax.mcmc.laplace_marginal.laplace_marginal_factory(
laplace_log_joint,
theta_init=jnp.zeros(n_theta),
)
# 2. Setup and run Window Adaptation for Laplace-HMC
start = time.time()
print("Running Laplace-HMC window adaptation on Funnel...")
warmup_laplace = blackjax.window_adaptation(
blackjax.laplace_hmc,
laplace_funnel,
num_integration_steps=3, # 3 leapfrog steps: each requires an L-BFGS solve,
# so fewer steps = much faster; 1D marginal mixes fine
)
key, warmup_key = jax.random.split(key)
(state_laplace, parameters_laplace), _ = warmup_laplace.run(warmup_key, jnp.array([0.0]), num_steps=500)
# 3. Sampling
sampler_laplace = blackjax.laplace_hmc(
laplace_log_joint,
theta_init=jnp.zeros(n_theta),
**parameters_laplace
)
print("Running Laplace-HMC sampling on Funnel...")
key, sample_key = jax.random.split(key)
states_laplace, _ = run_chains(sample_key, sampler_laplace.step, state_laplace, 400)
states_laplace.position.block_until_ready()
laplace_time = time.time() - start
# NUTS
start = time.time()
warmup = blackjax.window_adaptation(blackjax.nuts, nuts_log_joint)
key, warmup_key = jax.random.split(key)
(state_nuts, parameters_nuts), _ = warmup.run(warmup_key, {'v': jnp.array([0.0]), 'theta': jnp.zeros(n_theta)}, num_steps=1000)
nuts_sampler = blackjax.nuts(nuts_log_joint, **parameters_nuts)
print("Running NUTS on Funnel...")
key, sample_key = jax.random.split(key)
states_nuts, _ = run_chains(sample_key, nuts_sampler.step, state_nuts, 400)
states_nuts.position['v'].block_until_ready()
nuts_time = time.time() - start
print(f"Laplace-HMC time: {laplace_time:.2f}s")
print(f"NUTS time: {nuts_time:.2f}s")Running Laplace-HMC window adaptation on Funnel...
Running Laplace-HMC sampling on Funnel...
Running NUTS on Funnel...
Laplace-HMC time: 18.35s
NUTS time: 5.38s
from blackjax.diagnostics import effective_sample_size
fig, ax = plt.subplots(figsize=(8, 4))
v_laplace = states_laplace.position
v_nuts = states_nuts.position['v']
# Compute actual ESS (chains, draws, params=1)
ess_laplace = float(effective_sample_size(v_laplace[..., None]).sum())
ess_nuts = float(effective_sample_size(v_nuts[..., None]).sum())
ax.hist(np.array(v_laplace.flatten()), bins=30, density=True, alpha=0.6, color="steelblue",
label=f"Laplace-HMC ESS/min={60 * ess_laplace/laplace_time:.0f} ({laplace_time:.1f}s)")
ax.hist(np.array(v_nuts.flatten()), bins=30, density=True, alpha=0.5, color="orange",
label=f"NUTS ESS/min={60 * ess_nuts/nuts_time:.0f} ({nuts_time:.1f}s)")
# Exact marginal for v is N(0, 3^2)
v_range = jnp.linspace(-10, 10, 100)
ax.plot(v_range, jax.scipy.stats.norm.pdf(v_range, 0, 3), 'r--', label="Exact Marginal")
ax.set_title("Neal's Funnel: Marginal of v")
ax.legend()
plt.show()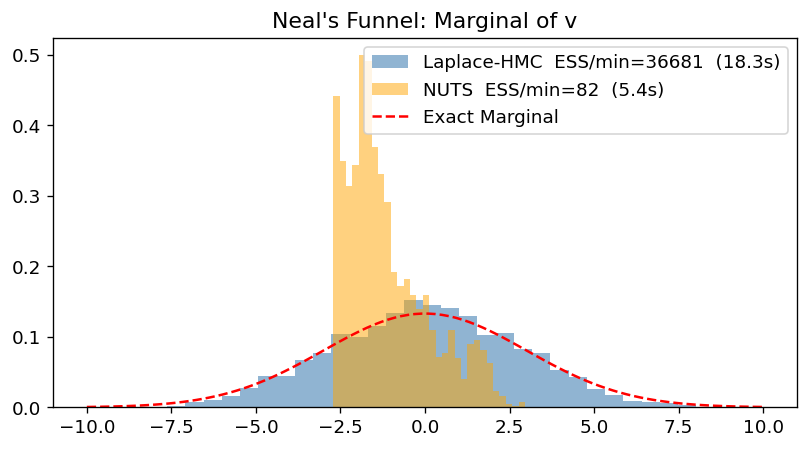
Section 2: Gaussian Process Regression (Laplace is Exact)¶
In a GP regression model with Gaussian likelihood the Laplace approximation is
exact: the conditional posterior is itself Gaussian, so
the marginal is computed without any approximation error.
Laplace-HMC samples the two kernel hyperparameters ;
the -dimensional function values are recovered afterwards via
laplace.sample_theta.
where is the RBF (squared-exponential) kernel. NUTS must explore the full -dimensional space whose geometry is funnel-shaped; Laplace-HMC only explores the 2-dimensional space.
# --- Data -----------------------------------------------------------------
n_gp = 30 # 30 training points
X_gp = jnp.linspace(0, 8, n_gp)
f_true = jnp.sin(X_gp)
sigma_n_gp = 0.3
key, k_data = jax.random.split(key)
y_gp = f_true + sigma_n_gp * jax.random.normal(k_data, (n_gp,))
# --- RBF kernel -----------------------------------------------------------
def rbf_kernel(x1, x2, log_ell, log_sigma_f):
ell, sigma_f = jnp.exp(log_ell), jnp.exp(log_sigma_f)
r2 = ((x1[:, None] - x2[None, :]) / ell) ** 2
return sigma_f ** 2 * jnp.exp(-0.5 * r2)
def log_joint_gp(theta, phi):
log_ell, log_sigma_f = phi[0], phi[1]
K = rbf_kernel(X_gp, X_gp, log_ell, log_sigma_f) + 1e-5 * jnp.eye(n_gp)
log_p_phi = (
jax.scipy.stats.norm.logpdf(log_ell, 0.0, 1.0)
+ jax.scipy.stats.norm.logpdf(log_sigma_f, 0.0, 1.0)
)
log_p_theta = jax.scipy.stats.multivariate_normal.logpdf(
theta, jnp.zeros(n_gp), K
)
log_lik = jax.scipy.stats.norm.logpdf(y_gp, theta, sigma_n_gp).sum()
return log_p_phi + log_p_theta + log_lik# --- NUTS on full joint (theta, phi) for comparison ----------------------
def nuts_log_joint_gp(params):
return log_joint_gp(params["theta"], params["phi"])
warmup_nuts_gp = blackjax.window_adaptation(blackjax.nuts, nuts_log_joint_gp)
key, warmup_key = jax.random.split(key)
(state_nuts_gp, params_nuts_gp), _ = warmup_nuts_gp.run(
warmup_key,
{"theta": jnp.zeros(n_gp), "phi": jnp.zeros(2)},
num_steps=500,
)
nuts_sampler_gp = blackjax.nuts(nuts_log_joint_gp, **params_nuts_gp)
n_nuts_gp = 400
print(f"Sampling full ({n_gp+2}-D) joint with NUTS ...")
key, sample_key = jax.random.split(key)
start = time.time()
states_nuts_gp, info = run_chains(
sample_key, nuts_sampler_gp.step, state_nuts_gp, n_nuts_gp
)
states_nuts_gp.position["theta"].block_until_ready()
nuts_gp_time = time.time() - start
phi_nuts_gp = states_nuts_gp.position["phi"]
print(f"Sampling full joint with NUTS done in {nuts_gp_time:.1f}s")Sampling full (32-D) joint with NUTS ...
Sampling full joint with NUTS done in 57.4s
# --- Laplace-HMC ----------------------------------------------------------
FIXED_STEPS = 4
laplace_gp = blackjax.mcmc.laplace_marginal.laplace_marginal_factory(
log_joint_gp, jnp.zeros(n_gp), maxiter=100
)
warmup_gp = blackjax.window_adaptation(
blackjax.laplace_mhmc, laplace_gp, num_integration_steps=FIXED_STEPS
)
key, warmup_key = jax.random.split(key)
(state_gp, params_gp), _ = warmup_gp.run(warmup_key, jnp.zeros(2), num_steps=500)
sampler_gp = blackjax.laplace_mhmc(log_joint_gp, theta_init=jnp.zeros(n_gp), **params_gp)
print("Sampling GP hyperparameters with Laplace-HMC ...")
key, sample_key = jax.random.split(key)
start = time.time()
states_gp, _ = run_chains(sample_key, sampler_gp.step, state_gp, 400)
states_gp.position.block_until_ready()
laplace_gp_time = time.time() - start
# Recover function values theta ~ N(theta_star, H^{-1}) for each phi sample
phi_gp_samples = states_gp.position
theta_star_samples = states_gp.theta_star
phi_flat = phi_gp_samples.reshape(-1, 2)
theta_star_flat = theta_star_samples.reshape(-1, n_gp)
key, rk = jax.random.split(key)
theta_gp_samples_flat = jax.vmap(laplace_gp.sample_theta)(
jax.random.split(rk, len(phi_flat)), phi_flat, theta_star_flat
)
theta_gp_samples = theta_gp_samples_flat.reshape(n_chains_default, -1, n_gp)
print(f"Sampling GP hyperparameters with Laplace-HMC done in {laplace_gp_time:.1f}s "
f"({phi_gp_samples.shape[0]}x{phi_gp_samples.shape[1]} samples)")Sampling GP hyperparameters with Laplace-HMC ...
Sampling GP hyperparameters with Laplace-HMC done in 43.8s (8x400 samples)
Source
theta_gp_mean = np.nanmean(theta_gp_samples, axis=(0, 1))
theta_gp_std = np.nanstd(theta_gp_samples, axis=(0, 1))
theta_nuts_mean = states_nuts_gp.position["theta"].mean((0, 1))
theta_nuts_std = states_nuts_gp.position["theta"].std((0, 1))
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
# Left: GP posterior at training points
ax = axes[0]
ax.scatter(np.array(X_gp), np.array(y_gp), s=20, alpha=0.5, color="gray", zorder=5, label="Data")
ax.plot(np.array(X_gp), np.array(f_true), "k--", lw=1.5, label="True f", zorder=4)
for mean, std, color, lbl in [
(theta_gp_mean, theta_gp_std, "steelblue", f"Laplace-HMC ({laplace_gp_time:.0f}s)"),
(theta_nuts_mean, theta_nuts_std, "orange",
f"NUTS on (θ,φ) ({nuts_gp_time:.0f}s)"),
]:
ax.plot(np.array(X_gp), np.array(mean), color=color, lw=2, label=lbl)
ax.fill_between(
np.array(X_gp),
np.array(mean - 2 * std),
np.array(mean + 2 * std),
alpha=0.2, color=color,
)
ax.set_xlabel("x")
ax.set_ylabel("f(x)")
ax.set_title("GP Posterior (mean ± 2σ)")
ax.legend(fontsize=9)
# Right: kernel hyperparameter posterior
ax = axes[1]
ax.scatter(
np.array(phi_gp_samples[..., 0].flatten()), np.array(phi_gp_samples[..., 1].flatten()),
s=8, alpha=0.3, color="steelblue", label="Laplace-HMC",
)
ax.scatter(
np.array(phi_nuts_gp[..., 0].flatten()), np.array(phi_nuts_gp[..., 1].flatten()),
s=8, alpha=0.3, color="orange", label="NUTS",
)
ax.set_xlabel("log length-scale (log ℓ)")
ax.set_ylabel("log amplitude (log σ_f)")
ax.set_title("Kernel Hyperparameter Posterior")
ax.legend(fontsize=9)
plt.tight_layout()
plt.show()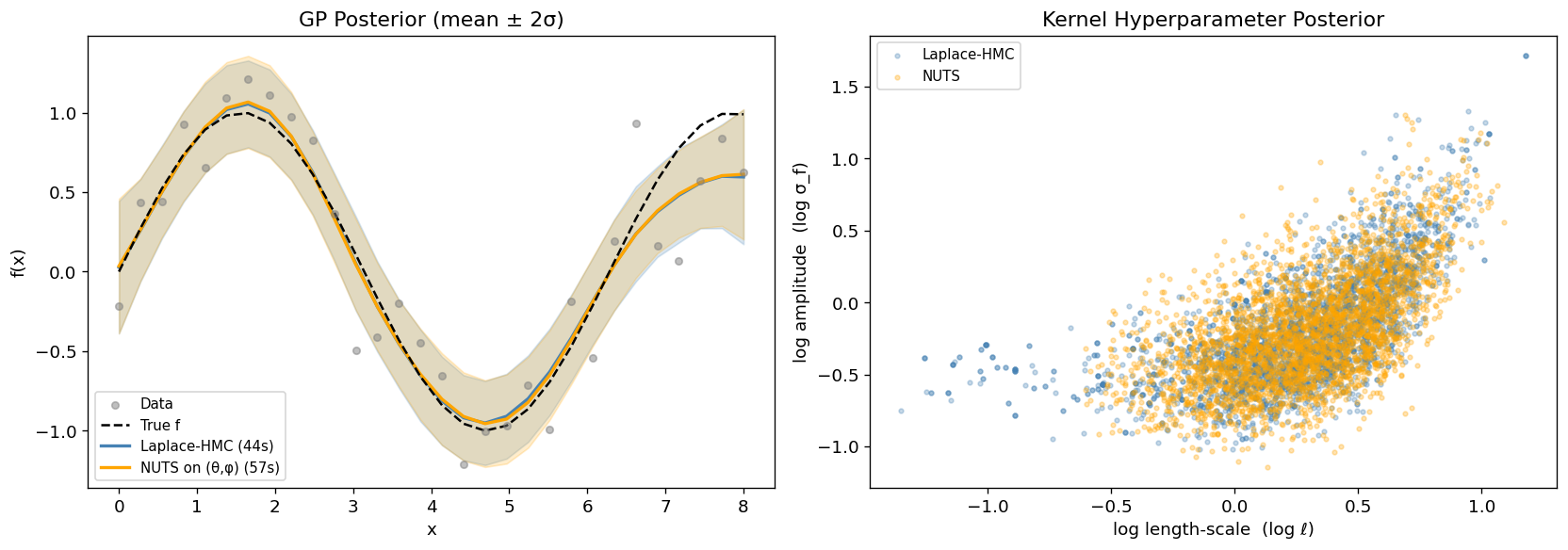
# Normalise by sample count for a fair per-sample comparison
time_per_sample_laplace = laplace_gp_time / (phi_gp_samples.shape[0] * phi_gp_samples.shape[1])
time_per_sample_nuts = nuts_gp_time / (phi_nuts_gp.shape[0] * phi_nuts_gp.shape[1])
speedup = time_per_sample_nuts / time_per_sample_laplace
print(f"Time / sample — Laplace-HMC: {time_per_sample_laplace*1000:.0f} ms "
f"NUTS: {time_per_sample_nuts*1000:.0f} ms ({speedup:.1f}x speedup)")
print(f"log length-scale: Laplace {float(phi_gp_samples[...,0].mean()):.2f} "
f"NUTS {float(phi_nuts_gp[...,0].mean()):.2f}")
print(f"log amplitude: Laplace {float(phi_gp_samples[...,1].mean()):.2f} "
f"NUTS {float(phi_nuts_gp[...,1].mean()):.2f}")Time / sample — Laplace-HMC: 14 ms NUTS: 18 ms (1.3x speedup)
log length-scale: Laplace 0.27 NUTS 0.28
log amplitude: Laplace -0.19 NUTS -0.20
Section 3: Comparing All Four Laplace-HMC Variants¶
The four top-level aliases form a 2×2 matrix along two independent axes:
| Endpoint + M-H | Multinomial trajectory | |
|---|---|---|
Fixed num_integration_steps | blackjax.laplace_hmc | blackjax.laplace_mhmc |
| Random step count per transition | blackjax.laplace_dhmc | blackjax.laplace_dmhmc |
Axis 1 — fixed vs. dynamic step count.
laplace_hmc and laplace_mhmc take a fixed num_integration_steps and plug
directly into window_adaptation.
laplace_dhmc and laplace_dmhmc draw the number of leapfrog steps uniformly at
random each transition, removing periodic-orbit sensitivity; they require a
rng_key argument at .init() and do not support window_adaptation.
Axis 2 — standard M-H vs. multinomial proposal.
Standard variants (hmc/dhmc) propose the trajectory endpoint and apply an M-H
accept/reject step; info.acceptance_rate is the usual Metropolis acceptance
probability (target ≈ 0.65).
Multinomial variants (mhmc/dmhmc) sample any trajectory point proportional to
exp(−energy) — is_accepted is always True, and info.acceptance_rate becomes
an average trajectory-weight diagnostic, not a reject probability.
We benchmark all four on the Gaussian Process Regression from Section 2
(30 latent variables, 2-D hyperparameter space φ = (log ℓ, log σ_f)).
The state_gp and params_gp already produced by Section 2’s
window_adaptation are reused directly — no extra warmup needed.
# ── Step 1: reuse the step_size / mass matrix tuned in Section 2 ─────────────
step_size_cmp = params_gp["step_size"]
inv_mass_cmp = params_gp["inverse_mass_matrix"]
phi_start = state_gp.position
print(f" step_size = {float(step_size_cmp):.4f}")
print(f" inv_mass_matrix = {np.array(inv_mass_cmp)}") step_size = 0.5009
inv_mass_matrix = [0.07190197 0.1309546 ]
# ── Step 2: build all four samplers with the same hyperparameters ─────────────
# Dynamic variants draw steps ~ Uniform[.5*FIXED_STEPS, 1.5*FIXED_STEPS], matching the mean
# trajectory length of the fixed-step variants.
integration_steps_fn = lambda k: jax.random.randint(k, (), int(.5 * FIXED_STEPS), int(1.5 * FIXED_STEPS))
theta0 = jnp.zeros(n_gp)
common_kwargs = dict(step_size=step_size_cmp, inverse_mass_matrix=inv_mass_cmp)
sampler_hmc = blackjax.laplace_hmc(
log_joint_gp, theta_init=theta0,
num_integration_steps=FIXED_STEPS, **common_kwargs,
)
sampler_mhmc = blackjax.laplace_mhmc(
log_joint_gp, theta_init=theta0,
num_integration_steps=FIXED_STEPS, **common_kwargs,
)
sampler_dhmc = blackjax.laplace_dhmc(
log_joint_gp, theta_init=theta0,
integration_steps_fn=integration_steps_fn, **common_kwargs,
)
sampler_dmhmc = blackjax.laplace_dmhmc(
log_joint_gp, theta_init=theta0,
integration_steps_fn=integration_steps_fn, **common_kwargs,
)# ── Step 3: initialise states ─────────────────────────────────────────────────
# Fixed-step variants: init(phi) → LaplaceHMCState
# Dynamic variants: init(phi, rng_key) → LaplaceDynamicHMCState
# rng_key seeds the per-step PRNG sequence
key, k1, k2 = jax.random.split(key, 3)
state_hmc = sampler_hmc.init(phi_start)
state_mhmc = sampler_mhmc.init(phi_start)
state_dhmc = sampler_dhmc.init(phi_start, k1) # extra rng_key required
state_dmhmc = sampler_dmhmc.init(phi_start, k2) # extra rng_key required
print("LaplaceHMCState fields: ", list(state_hmc._fields))
print("LaplaceDynamicHMCState fields: ", list(state_dhmc._fields))LaplaceHMCState fields: ['position', 'logdensity', 'logdensity_grad', 'theta_star']
LaplaceDynamicHMCState fields: ['position', 'logdensity', 'logdensity_grad', 'theta_star', 'random_generator_arg']
# ── Step 4: sample and compare ───────────────────────────────────────────────
N_SAMPLES = 400
cmp_results = {}
for name, sampler, state in [
("laplace_hmc", sampler_hmc, state_hmc),
("laplace_mhmc", sampler_mhmc, state_mhmc),
("laplace_dhmc", sampler_dhmc, state_dhmc),
("laplace_dmhmc", sampler_dmhmc, state_dmhmc),
]:
key, sample_key = jax.random.split(key)
t0 = time.time()
states, info = run_chains(sample_key, sampler.step, state, N_SAMPLES)
states.position.block_until_ready()
elapsed = time.time() - t0
phi_post = states.position
ess_per_param = effective_sample_size(phi_post) # (chains, draws, params)
ess_min = float(jnp.min(ess_per_param))
# acceptance_rate semantics differ by proposal type:
# hmc / dhmc → M-H probability (target ≈ 0.65)
# mhmc / dmhmc → avg trajectory weight (diagnostic; is_accepted always True)
is_mh = name in ("laplace_hmc", "laplace_dhmc")
acc_mean = float(info.acceptance_rate.mean())
acc_label = "M-H acc" if is_mh else "traj weight"
cmp_results[name] = dict(ess=ess_min, time=elapsed,
accept=acc_mean, is_mh=is_mh, acc_label=acc_label)
print(
f"{name:20s} ESS={ess_min:5.0f} time={elapsed:.1f}s "
f"ESS/min={60 * ess_min/elapsed:5.0f} {acc_label}={acc_mean:.2f}"
)laplace_hmc ESS= 47 time=33.3s ESS/min= 85 M-H acc=0.70
laplace_mhmc ESS= 349 time=38.6s ESS/min= 542 traj weight=0.84
laplace_dhmc ESS= 43 time=35.8s ESS/min= 72 M-H acc=0.71
laplace_dmhmc ESS= 385 time=38.6s ESS/min= 598 traj weight=0.86
Source
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
names = list(cmp_results.keys())
colors = ["#4878CF", "#6ACC65", "#D65F5F", "#B47CC7"]
axes[0].bar(names, [cmp_results[n]["ess"] for n in names], color=colors)
axes[0].set_ylabel("Min ESS")
axes[0].set_title("Effective Sample Size")
axes[0].tick_params(axis="x", rotation=18)
axes[1].bar(names, [60 * cmp_results[n]["ess"] / cmp_results[n]["time"] for n in names], color=colors)
axes[1].set_ylabel("Min ESS / minute")
axes[1].set_title("Sampling Efficiency")
axes[1].tick_params(axis="x", rotation=18)
ax = axes[2]
for i, n in enumerate(names):
r = cmp_results[n]
ax.bar(i, r["accept"], color=colors[i])
ax.text(i, r["accept"] + 0.02, r["acc_label"],
ha="center", va="bottom", fontsize=7)
ax.axhline(0.65, ls="--", color="gray", lw=1, label="0.65 M-H target")
ax.set_ylim(0, 1.35)
ax.set_xticks(range(len(names)))
ax.set_xticklabels(names, rotation=18)
ax.set_ylabel("Mean value")
ax.set_title("Acceptance / Trajectory Weight")
ax.legend(fontsize=8)
plt.suptitle(
"GP Regression — four Laplace-HMC variants\n"
f"(shared step_size={float(step_size_cmp):.3f}, "
f"mean trajectory length={FIXED_STEPS} leapfrog steps)",
fontsize=10,
)
plt.tight_layout()
plt.show()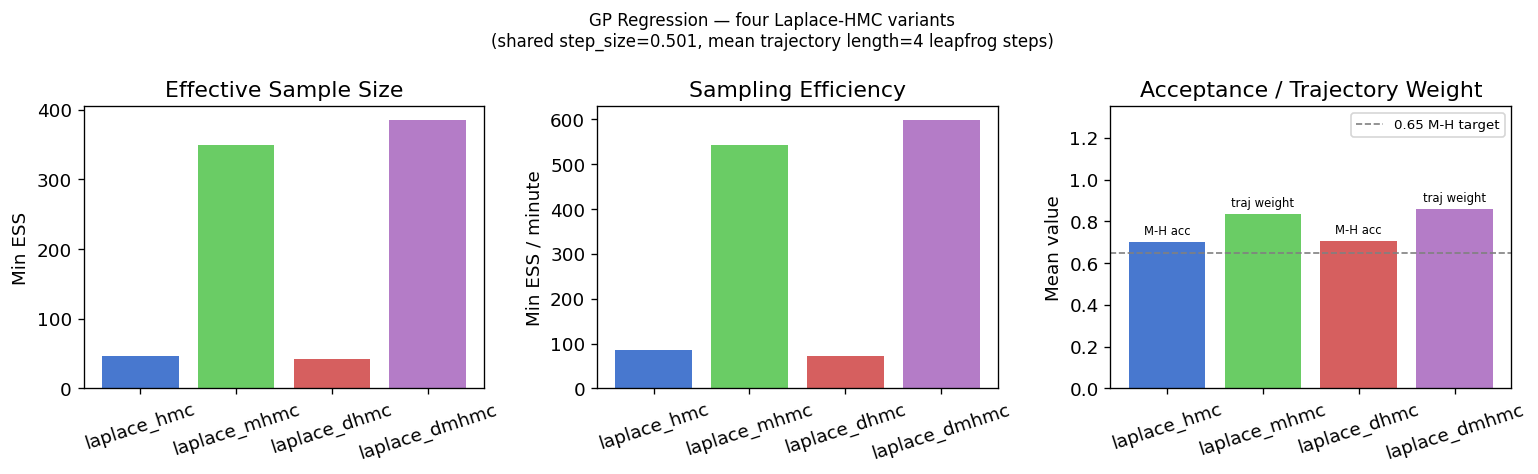
Reading the results.
The acceptance rate panel reveals something important: laplace_hmc can show acceptable M-H
acceptance rate yet produce a very low ESS.
This could be the classic periodic-orbit pathology — a fixed trajectory length that
happens to nearly return to the starting point, causing the chain to take
tiny effective steps despite technically accepting.
laplace_dhmc breaks the orbit by randomising the step count each transition, so
even the same step size yields much better mixing.
Multinomial variants (laplace_mhmc, laplace_dmhmc) also avoid the trap: by
sampling a random point along the trajectory they implicitly vary the effective
displacement, and is_accepted is always True so they never waste a trajectory.
This illustrates the core trade-off: if you can tune num_integration_steps
carefully, laplace_hmc is the cheapest option; if you are unsure of the right
trajectory length (common in practice), the dynamic and multinomial variants are
more robust out of the box.
Quick-reference: when to use each variant.
| Variant | Use when |
|---|---|
laplace_hmc | Default when trajectory length is well-tuned; window_adaptation works directly |
laplace_mhmc | Drop-in upgrade when M-H acceptance is low or ESS per gradient is poor |
laplace_dhmc | Trajectory length is hard to tune; randomised steps break periodic-orbit traps |
laplace_dmhmc | Combines both benefits — best robustness for unknown geometry |