Multimodal distributions are typically hard to sample from, in particular using energy based methods such as HMC, as you need high energy levels to escape a potential well.
Tempered SMC helps with this by considering a sequence of distributions:
where the tempering parameter takes increasing values between 0 and 1. Tempered SMC will also particularly shine when the MCMC step is not well calibrated (too small step size, etc) like in the example below.
Imports¶
Notebook Cell
import matplotlib.pyplot as plt
plt.rcParams["axes.spines.right"] = False
plt.rcParams["axes.spines.top"] = Falseimport jax
from datetime import date
rng_key = jax.random.key(int(date.today().strftime("%Y%m%d")))import numpy as np
import jax.numpy as jnp
from jax.scipy.stats import multivariate_normal
import blackjax
import blackjax.smc.resampling as resampling
from blackjax.smc import extend_paramsSampling From a Bimodal Potential¶
Experimental Setup¶
We consider a prior distribution
and a potential function
This corresponds to the following distribution. We plot the resulting tempered density for 5 different values of : from which correponds to the original density to . The lower the value of the easier it is for the sampler to jump between the modes of the posterior density.
def V(x):
return 5 * jnp.square(jnp.sum(x**2, axis=-1) - 1)
def prior_log_prob(x):
d = x.shape[-1]
return multivariate_normal.logpdf(x, jnp.zeros((d,)), jnp.eye(d))
linspace = jnp.linspace(-2, 2, 5000)[..., None]
lambdas = jnp.linspace(0.0, 1.0, 5)
prior_logvals = prior_log_prob(linspace)
potential_vals = V(linspace)
log_res = prior_logvals - lambdas[..., None] * potential_vals
density = jnp.exp(log_res)
normalizing_factor = jnp.sum(density, axis=1, keepdims=True) * (
linspace[1] - linspace[0]
)
density /= normalizing_factorSource
fig, ax = plt.subplots(figsize=(12, 8))
ax.plot(linspace.squeeze(), density.T)
ax.legend(list(lambdas))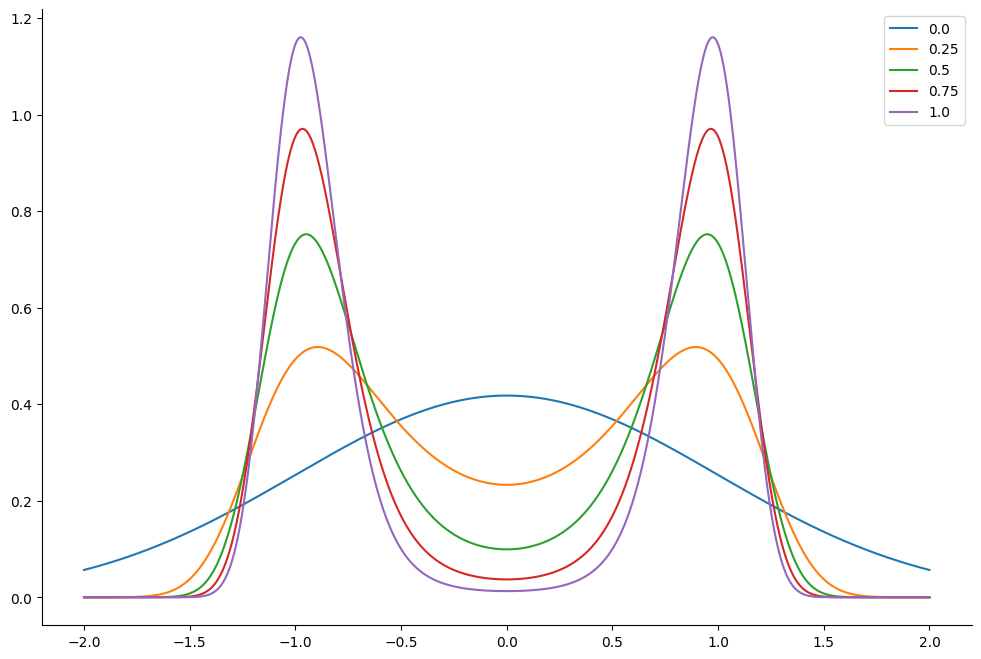
def inference_loop(rng_key, mcmc_kernel, initial_state, num_samples):
@jax.jit
def one_step(state, k):
state, _ = mcmc_kernel(k, state)
return state, state
keys = jax.random.split(rng_key, num_samples)
_, states = jax.lax.scan(one_step, initial_state, keys)
return states
def full_logdensity(x):
return -V(x) + prior_log_prob(x)
inv_mass_matrix = jnp.eye(1)
n_samples = 10_000Sample with HMC¶
We first try to sample from the posterior density using an HMC kernel.
%%time
hmc_parameters = dict(
step_size=1e-4, inverse_mass_matrix=inv_mass_matrix, num_integration_steps=50
)
hmc = blackjax.hmc(full_logdensity, **hmc_parameters)
hmc_state = hmc.init(jnp.ones((1,)))
rng_key, sample_key = jax.random.split(rng_key)
hmc_samples = inference_loop(sample_key, hmc.step, hmc_state, n_samples)CPU times: user 3.51 s, sys: 173 ms, total: 3.68 s
Wall time: 2.62 s
Source
samples = np.array(hmc_samples.position[:, 0])
_ = plt.hist(samples, bins=100, density=True)
_ = plt.plot(linspace.squeeze(), density[-1])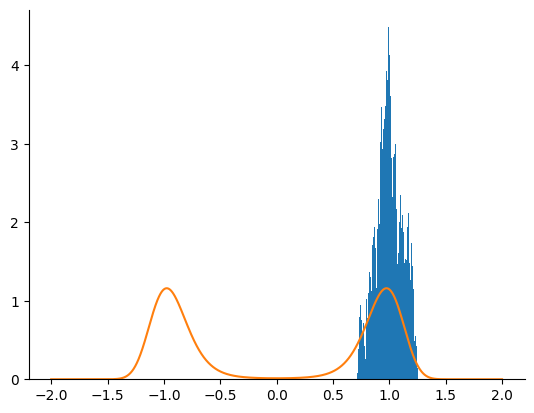
Sample with NUTS¶
We now use a NUTS kernel.
%%time
nuts_parameters = dict(step_size=1e-4, inverse_mass_matrix=inv_mass_matrix)
nuts = blackjax.nuts(full_logdensity, **nuts_parameters)
nuts_state = nuts.init(jnp.ones((1,)))
rng_key, sample_key = jax.random.split(rng_key)
nuts_samples = inference_loop(sample_key, nuts.step, nuts_state, n_samples)CPU times: user 26.3 s, sys: 253 ms, total: 26.6 s
Wall time: 25.5 s
Source
samples = np.array(nuts_samples.position[:, 0])
_ = plt.hist(samples, bins=100, density=True)
_ = plt.plot(linspace.squeeze(), density[-1])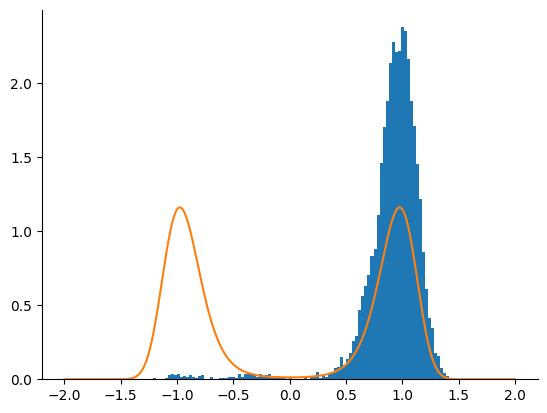
Tempered SMC with HMC Kernel¶
We now use the adaptive tempered SMC algorithm with an HMC kernel. We only take one HMC step before resampling. The algorithm is run until crosses the limit.
def smc_inference_loop(rng_key, smc_kernel, initial_state):
"""Run the temepered SMC algorithm.
We run the adaptive algorithm until the tempering parameter lambda reaches the value
lambda=1.
"""
def cond(carry):
i, state, _k = carry
return state.tempering_param < 1
def one_step(carry):
i, state, k = carry
k, subk = jax.random.split(k, 2)
state, _ = smc_kernel(subk, state)
return i + 1, state, k
n_iter, final_state, _ = jax.lax.while_loop(
cond, one_step, (0, initial_state, rng_key)
)
return n_iter, final_state%%time
loglikelihood = lambda x: -V(x)
hmc_parameters = dict(
step_size=1e-4, inverse_mass_matrix=inv_mass_matrix, num_integration_steps=1
)
tempered = blackjax.adaptive_tempered_smc(
prior_log_prob,
loglikelihood,
blackjax.hmc.build_kernel(),
blackjax.hmc.init,
extend_params(hmc_parameters),
resampling.systematic,
0.5,
num_mcmc_steps=1,
)
rng_key, init_key, sample_key = jax.random.split(rng_key, 3)
initial_smc_state = jax.random.multivariate_normal(
init_key, jnp.zeros([1]), jnp.eye(1), (n_samples,)
)
initial_smc_state = tempered.init(initial_smc_state)
n_iter, smc_samples = smc_inference_loop(sample_key, tempered.step, initial_smc_state)
print("Number of steps in the adaptive algorithm: ", n_iter.item())Number of steps in the adaptive algorithm: 13
CPU times: user 5.59 s, sys: 201 ms, total: 5.79 s
Wall time: 3.26 s
Source
samples = np.array(smc_samples.particles[:, 0])
_ = plt.hist(samples, bins=100, density=True)
_ = plt.plot(linspace.squeeze(), density[-1])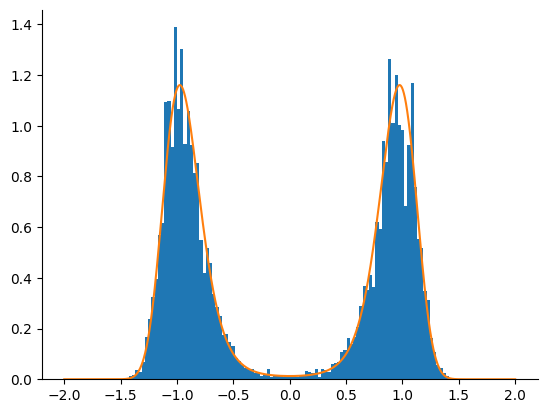
Sampling from the Rastrigin Potential¶
Experimental Setup¶
We consider a prior distribution and we want to sample from a Rastrigin type potential function where we choose . These potential functions are known to be particularly hard to sample.
We plot the resulting tempered density for 5 different values of : from which correponds to the original density to . The lower the value of the easier it is to sampler from the posterior log-density.
def V(x):
d = x.shape[-1]
res = -10 * d + jnp.sum(x**2 - 10 * jnp.cos(2 * jnp.pi * x), -1)
return res
linspace = jnp.linspace(-5, 5, 5000)[..., None]
lambdas = jnp.linspace(0.0, 1.0, 5)
potential_vals = V(linspace)
log_res = lambdas[..., None] * potential_vals
density = jnp.exp(-log_res)
normalizing_factor = jnp.sum(density, axis=1, keepdims=True) * (
linspace[1] - linspace[0]
)
density /= normalizing_factorSource
fig, ax = plt.subplots(figsize=(12, 8))
ax.semilogy(linspace.squeeze(), density.T)
ax.legend(list(lambdas))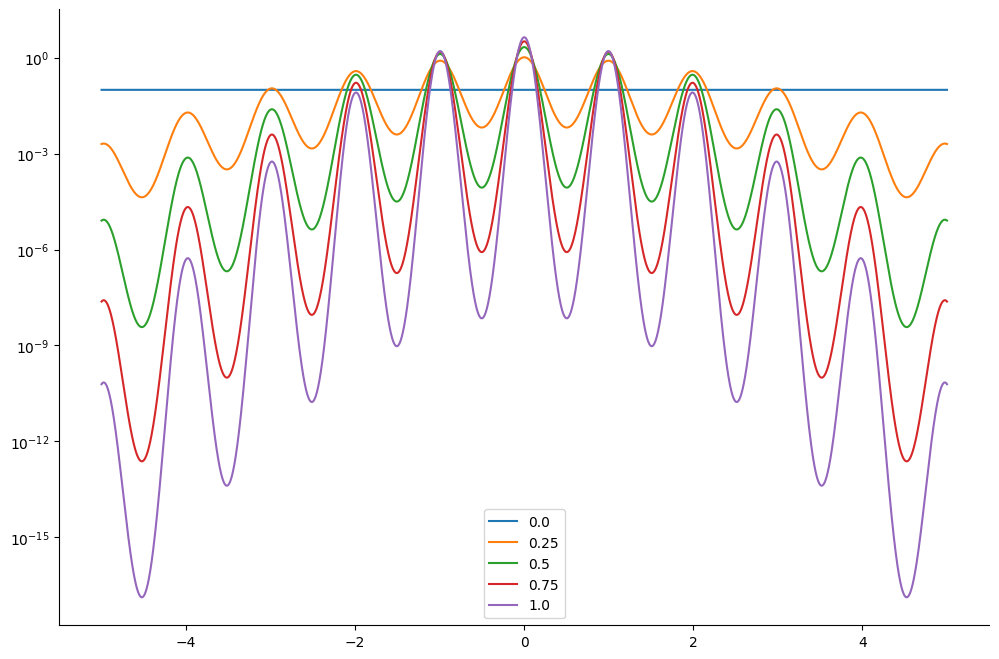
def inference_loop(rng_key, mcmc_kernel, initial_state, num_samples):
def one_step(state, k):
state, _ = mcmc_kernel(k, state)
return state, state
keys = jax.random.split(rng_key, num_samples)
_, states = jax.lax.scan(one_step, initial_state, keys)
return states
inv_mass_matrix = jnp.eye(1)
n_samples = 1_000HMC Sampler¶
We first try to sample from the posterior density using an HMC kernel.
%%time
loglikelihood = lambda x: -V(x)
hmc_parameters = dict(
step_size=1e-2, inverse_mass_matrix=inv_mass_matrix, num_integration_steps=50
)
hmc = blackjax.hmc(full_logdensity, **hmc_parameters)
hmc_state = hmc.init(jnp.ones((1,)))
rng_key, sample_key = jax.random.split(rng_key)
hmc_samples = inference_loop(sample_key, hmc.step, hmc_state, n_samples)CPU times: user 1.66 s, sys: 41.7 ms, total: 1.7 s
Wall time: 1.16 s
Source
samples = np.array(hmc_samples.position[:, 0])
_ = plt.hist(samples, bins=100, density=True)
_ = plt.plot(linspace.squeeze(), density[-1])
_ = plt.yscale("log")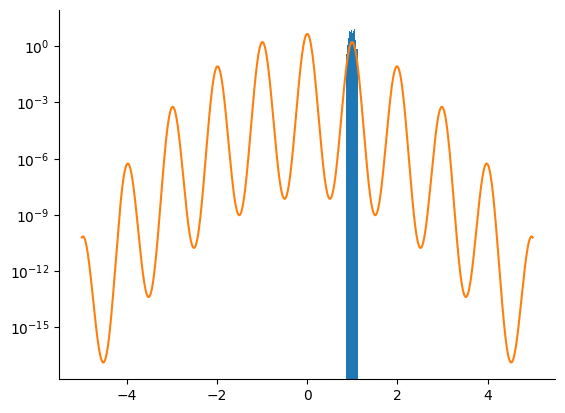
NUTS Sampler¶
We do the same using a NUTS kernel.
%%time
nuts_parameters = dict(step_size=1e-2, inverse_mass_matrix=inv_mass_matrix)
nuts = blackjax.nuts(full_logdensity, **nuts_parameters)
nuts_state = nuts.init(jnp.ones((1,)))
rng_key, sample_key = jax.random.split(rng_key)
nuts_samples = inference_loop(sample_key, nuts.step, nuts_state, n_samples)CPU times: user 1.97 s, sys: 31.1 ms, total: 2 s
Wall time: 1.22 s
Source
samples = np.array(nuts_samples.position[:, 0])
_ = plt.hist(samples, bins=100, density=True)
_ = plt.plot(linspace.squeeze(), density[-1])
_ = plt.yscale("log")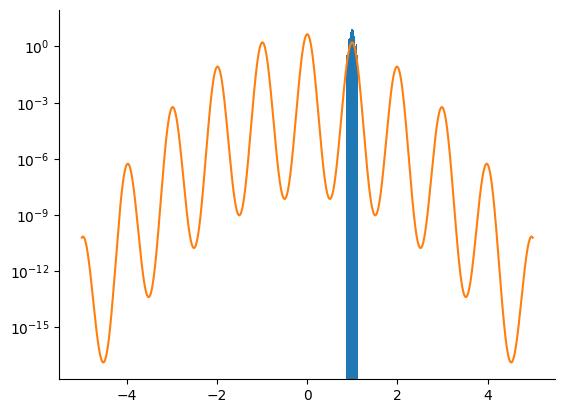
Tempered SMC with HMC Kernel¶
We now use the adaptive tempered SMC algorithm with an HMC kernel. We only take one HMC step before resampling. The algorithm is run until crosses the limit. We correct the bias introduced by the (arbitrary) prior.
%%time
loglikelihood = lambda x: -V(x)
hmc_parameters = dict(
step_size=1e-2, inverse_mass_matrix=inv_mass_matrix, num_integration_steps=100
)
tempered = blackjax.adaptive_tempered_smc(
prior_log_prob,
loglikelihood,
blackjax.hmc.build_kernel(),
blackjax.hmc.init,
extend_params(hmc_parameters),
resampling.systematic,
0.75,
num_mcmc_steps=1,
)
rng_key, init_key, sample_key = jax.random.split(rng_key, 3)
initial_smc_state = jax.random.multivariate_normal(
init_key, jnp.zeros([1]), jnp.eye(1), (n_samples,)
)
initial_smc_state = tempered.init(initial_smc_state)
n_iter, smc_samples = smc_inference_loop(sample_key, tempered.step, initial_smc_state)
print("Number of steps in the adaptive algorithm: ", n_iter.item())Number of steps in the adaptive algorithm: 9
CPU times: user 4.36 s, sys: 129 ms, total: 4.49 s
Wall time: 2.47 s
Source
samples = np.array(smc_samples.particles[:, 0])
_ = plt.hist(samples, bins=100, density=True)
_ = plt.plot(linspace.squeeze(), density[-1])
_ = plt.yscale("log")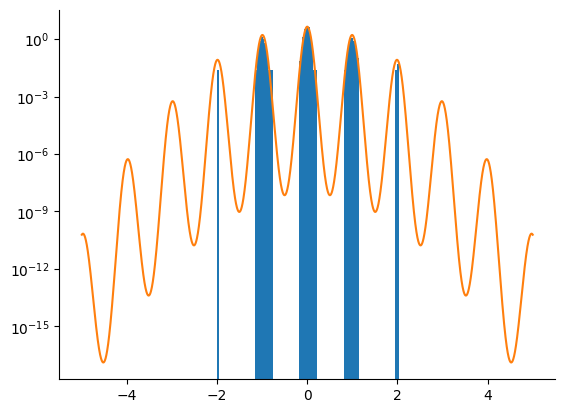
The tempered SMC algorithm with the HMC kernel clearly outperfoms the HMC and NUTS kernels alone.