This example replicates the case study analyzing financial time series, specifically the daily difference in log price data of Google’s stock, referred to as returns .
We’ll assume that at any given time the stock’s returns will follow one of two regimes: an independent random walk regime where and an autoregressive regime where . Being on either of the two regimes, , will depend on the previous time’s regime , call these probabilities for . Set as parameters of the model and and define the complementary probabilities by definition: and . Since the regime at any time is unobserved, we instead carry over time the probability of belonging to either one regime as . Finally, we need to model initial values, both for returns and probability of belonging to one of the two regimes .
In the whole, our regime-switching model is defined by the likelihood
where , , and for . And the priors of the parameters are:
where indicates the truncated at 0 Gaussian distribution and the half-Cauchy distribution.
Notebook Cell
import matplotlib.pyplot as plt
import arviz as az
plt.rcParams["axes.spines.right"] = False
plt.rcParams["axes.spines.top"] = False
az.rcParams["plot.max_subplots"] = 200import jax
jax.config.update("jax_enable_x64", True)
from datetime import date
rng_key = jax.random.key(int(date.today().strftime("%Y%m%d")))import jax.numpy as jnp
import numpy as np
import numpyro
import numpyro.distributions as distrib
import pandas as pd
from jax.scipy.stats import norm
from numpyro.diagnostics import print_summary
from numpyro.infer.util import initialize_model
import blackjax
class RegimeMixtureDistribution(distrib.Distribution):
arg_constraints = {
"alpha": distrib.constraints.real,
"rho": distrib.constraints.positive,
"sigma": distrib.constraints.positive,
"p": distrib.constraints.interval(0, 1),
"xi_0": distrib.constraints.interval(0, 1),
"y_0": distrib.constraints.real,
"T": distrib.constraints.positive_integer,
}
support = distrib.constraints.real
def __init__(self, alpha, rho, sigma, p, xi_0, y_0, T, validate_args=True):
self.alpha, self.rho, self.sigma, self.p, self.xi_0, self.y_0, self.T = (
alpha,
rho,
sigma,
p,
xi_0,
y_0,
T,
)
super().__init__(event_shape=(T,), validate_args=validate_args)
def log_prob(self, value):
def obs_t(carry, y):
y_prev, log_xi = carry # log_xi: [log P(s_{t-1}=1), log P(s_{t-1}=2)]
log_eta_1 = norm.logpdf(y, loc=self.alpha[0], scale=self.sigma[0])
log_eta_2 = norm.logpdf(
y, loc=self.alpha[1] + y_prev * self.rho, scale=self.sigma[1]
)
# log P(y_t | s_{t-1} = j) for j in {1, 2}
log_lik_1 = jnp.logaddexp(
jnp.log(self.p[0]) + log_eta_1,
jnp.log1p(-self.p[0]) + log_eta_2,
)
log_lik_2 = jnp.logaddexp(
jnp.log1p(-self.p[1]) + log_eta_1,
jnp.log(self.p[1]) + log_eta_2,
)
log_liks = jnp.array([log_lik_1, log_lik_2])
log_xi_unnorm = log_xi + log_liks
log_lik_total = jax.nn.logsumexp(log_xi_unnorm)
new_log_xi = log_xi_unnorm - log_lik_total
return (y, new_log_xi), log_lik_total
log_xi_0 = jnp.log(jnp.array([self.xi_0, 1.0 - self.xi_0]))
_, log_liks = jax.lax.scan(obs_t, (self.y_0, log_xi_0), value)
return jnp.sum(log_liks)
def sample(self, key, sample_shape=()):
return jnp.zeros(sample_shape + self.event_shape)
class RegimeSwitchHMM:
def __init__(self, T, y) -> None:
self.T = T
self.y = y
def model(self, y=None):
rho = numpyro.sample("rho", distrib.TruncatedNormal(1.0, 0.1, low=0.0))
alpha = numpyro.sample("alpha", distrib.Normal(0.0, 0.1).expand([2]))
sigma = numpyro.sample("sigma", distrib.HalfCauchy(1.0).expand([2]))
p = numpyro.sample("p", distrib.Beta(10.0, 2.0).expand([2]))
xi_0 = numpyro.sample("xi_0", distrib.Beta(2.0, 2.0))
y_0 = numpyro.sample("y_0", distrib.Normal(0.0, 0.1))
numpyro.sample(
"obs",
RegimeMixtureDistribution(alpha, rho, sigma, p, xi_0, y_0, self.T),
obs=y,
)
def initialize_model(self, rng_key, n_chain):
(init_params, *_), self.potential_fn, *_ = initialize_model(
rng_key,
self.model,
model_kwargs={"y": self.y},
dynamic_args=True,
)
# Separate the two regimes by anchoring sigma at [3, 10] in constrained
# space (numpyro uses log transform, so unconstrained = log(constrained)).
# Without this, chains near sigma[0] ≈ sigma[1] can fall into degenerate
# modes where one regime becomes inactive.
init_params = dict(init_params)
init_params["sigma"] = jnp.log(jnp.array([3.0, 10.0]))
init_params["rho"] = jnp.zeros(()) # log(1) = 0, the prior mode
flat, unravel_fn = jax.flatten_util.ravel_pytree(init_params)
kchain = jax.random.split(rng_key, n_chain)
self.init_params = jax.vmap(
lambda k: unravel_fn(flat + 0.1 * jax.random.normal(k, flat.shape))
)(kchain)
def logdensity_fn(self, params):
return -self.potential_fn(self.y)(params)
def inference_loop(rng, init_state, kernel, n_iter):
keys = jax.random.split(rng, n_iter)
def step(state, key):
state, info = kernel(key, state)
return state, (state, info)
_, (states, info) = jax.lax.scan(step, init_state, keys)
return states, infourl = "https://raw.githubusercontent.com/blackjax-devs/blackjax/main/docs/examples/data/google.csv"
data = pd.read_csv(url)
y = data["dl_ac"].values * 100
T, _ = data.shapedist = RegimeSwitchHMM(T, y)n_chain, n_warm, n_iter = 8, 2000, 2000
ksam, kinit = jax.random.split(jax.random.key(0), 2)
dist.initialize_model(kinit, n_chain)tic1 = pd.Timestamp.now()
k_warm, k_sample = jax.random.split(ksam)
(_, parameters), _ = blackjax.window_adaptation(
blackjax.nuts, dist.logdensity_fn
).run(k_warm, jax.tree.map(lambda x: x[0], dist.init_params), n_warm)
kernel = blackjax.nuts(dist.logdensity_fn, **parameters).step
def one_chain(k_sam, init_param):
init_state = blackjax.nuts(dist.logdensity_fn, **parameters).init(init_param)
state, info = inference_loop(k_sam, init_state, kernel, n_iter)
return state.position, info
k_sample = jax.random.split(k_sample, n_chain)
samples, infos = jax.vmap(one_chain)(k_sample, dist.init_params)
tic2 = pd.Timestamp.now()
print("Runtime for NUTS", tic2 - tic1)Runtime for NUTS 0 days 00:01:32.570961
print_summary(samples)
mean std median 5.0% 95.0% n_eff r_hat
alpha[0] 0.06 0.08 0.06 -0.08 0.20 27827.21 1.00
alpha[1] 0.01 0.10 0.01 -0.15 0.18 26813.28 1.00
p[0] 2.71 0.42 2.69 2.05 3.37 6837.55 1.00
p[1] 1.82 0.91 1.75 0.46 3.26 10453.65 1.00
rho -0.02 0.11 -0.01 -0.18 0.16 26029.82 1.00
sigma[0] 1.04 0.05 1.04 0.97 1.12 21694.94 1.00
sigma[1] 2.16 0.18 2.14 1.85 2.44 16638.53 1.00
xi_0 0.50 1.04 0.47 -1.18 2.24 25371.45 1.00
y_0 0.00 0.10 0.00 -0.16 0.17 25656.75 1.00
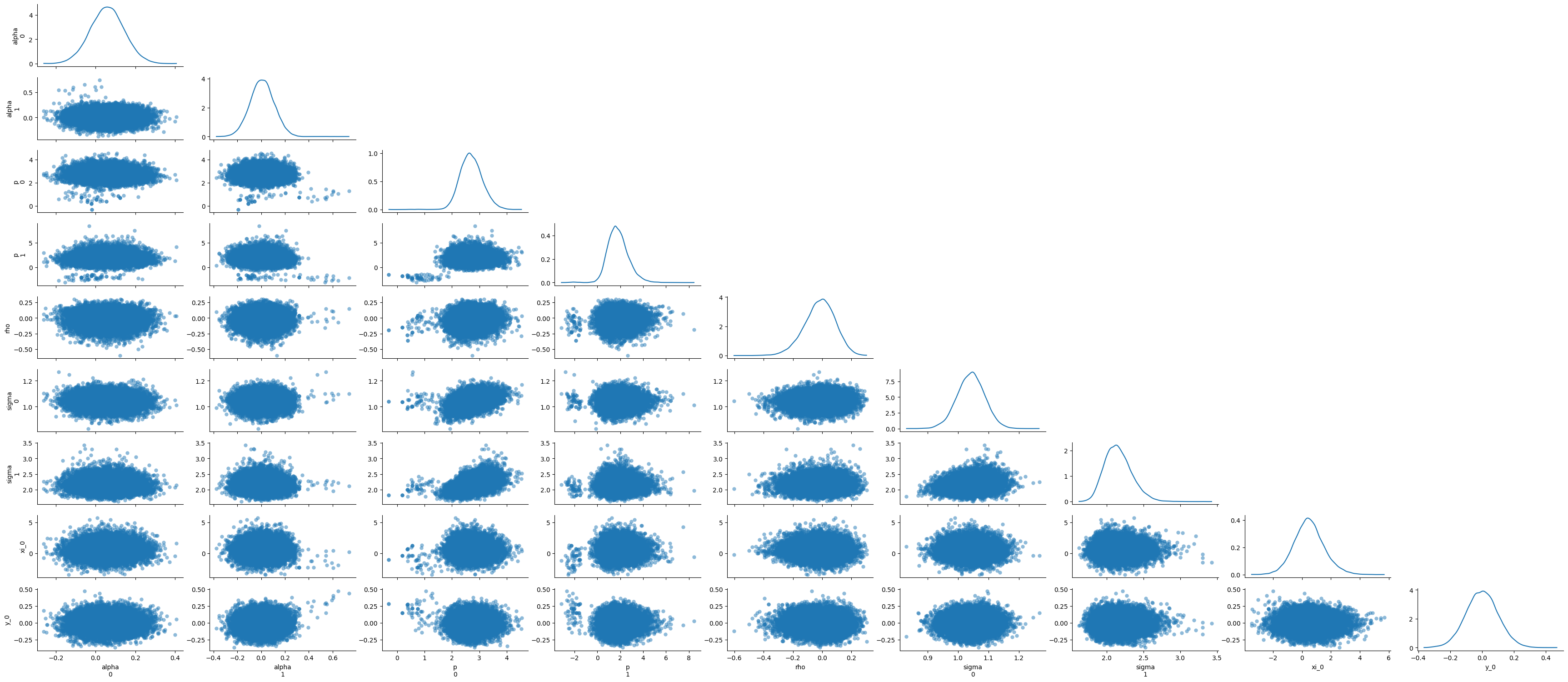
idata = az.from_dict({"posterior": samples})
az.plot_pair(idata, marginal=True, marginal_kind='kde')
plt.tight_layout();