Use with PyTensor models#
Blackjax accepts any log-probability function as long as it is compatible with jax.jit,
jax.grad (for gradient-based samplers) and jax.vmap. In this example we show how to
use PyTensor as a modeling language and Blackjax as an
inference library.
```{note}
This notebook used to demonstrate Aesara integration.
Aesara has been archived and is incompatible with NumPy 2.x.
PyTensor is its actively-maintained successor with an almost identical API.
We implement the following Binomial response model for the rat tumor dataset:
$$ \begin{align*} Y &\sim \operatorname{Binomial}(N, \theta)\ \theta &\sim \operatorname{Beta}(\alpha, \beta)\ \alpha, \beta &\sim \frac{1}{(\alpha + \beta)^{2.5}} \end{align*} $$
We sample in the unconstrained space: \(\log\alpha\), \(\log\beta\), and \(\text{logit}(\theta)\).
import jax
from datetime import date
rng_key = jax.random.key(int(date.today().strftime("%Y%m%d")))
We build the log-density graph symbolically in PyTensor. We work in the unconstrained space using \(\log\alpha\), \(\log\beta\), and \(\text{logit}(\theta_i)\) as free parameters and include the log-Jacobian corrections for the transforms:
import pytensor
import pytensor.tensor as pt
from pytensor.tensor.special import gammaln
log_a = pt.scalar('log_a')
log_b = pt.scalar('log_b')
logit_theta = pt.vector('logit_theta')
a = pt.exp(log_a)
b = pt.exp(log_b)
theta = pt.sigmoid(logit_theta)
# Improper prior: -2.5 * log(alpha + beta)
logprior_ab = -2.5 * pt.log(a + b)
# Log-Jacobians of the parameter transforms
logdet_a = log_a
logdet_b = log_b
logdet_theta = pt.sum(pt.log(theta) + pt.log(1 - theta))
# Beta(a, b) log-density for each theta_i
logp_beta = pt.sum(
gammaln(a + b) - gammaln(a) - gammaln(b)
+ (a - 1) * pt.log(theta)
+ (b - 1) * pt.log(1 - theta)
)
# Binomial log-likelihood
logp_binom = pt.sum(
n_of_positives * pt.log(theta)
+ (group_size - n_of_positives) * pt.log(1 - theta)
)
logdensity = logprior_ab + logdet_a + logdet_b + logdet_theta + logp_beta + logp_binom
To sample with Blackjax we compile the log-density graph using PyTensor’s JAX backend,
which produces a function fully compatible with jax.jit and jax.grad:
fn_jax = pytensor.function([log_a, log_b, logit_theta], logdensity, mode="JAX")
jit_fn = fn_jax.vm.jit_fn # the underlying JAX function
def logdensity_fn(position):
"""Wrap positional args into a dict-compatible interface for Blackjax."""
return jit_fn(position["log_a"], position["log_b"], position["logit_theta"])[0]
Let’s define the initial position and run window adaptation for the NUTS sampler:
import blackjax
def init_param_fn(seed):
key1, key2, key3 = jax.random.split(seed, 3)
return {
"log_a": jax.random.normal(key1, ()),
"log_b": jax.random.normal(key2, ()),
"logit_theta": jax.random.normal(key3, (n_rat_tumors,)),
}
rng_key, init_key, warmup_key, sample_key = jax.random.split(rng_key, 4)
init_position = init_param_fn(init_key)
adapt = blackjax.window_adaptation(blackjax.nuts, logdensity_fn)
(state, parameters), _ = adapt.run(warmup_key, init_position, 1000)
kernel = blackjax.nuts(logdensity_fn, **parameters).step
Now we run the inference loop:
states, infos = inference_loop(sample_key, kernel, state, 1000)
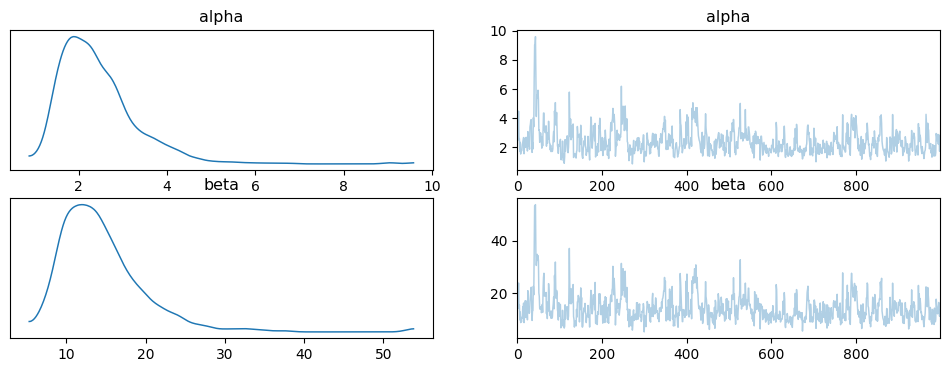
And plot the posterior samples on the original (constrained) scale using ArviZ:
import numpy as np
import arviz as az
posterior = {
"alpha": np.exp(states.position["log_a"]),
"beta": np.exp(states.position["log_b"]),
}
idata = az.from_dict(posterior={k: v[None, ...] for k, v in posterior.items()})
az.plot_trace(idata);